PROTEIN ENGINEERING
prof. Mgr. Jiří Damborský, Dr.
E-mail: jiri.damborsky@fnusa.cz
We educate leaders for Science and Business!
Research focus
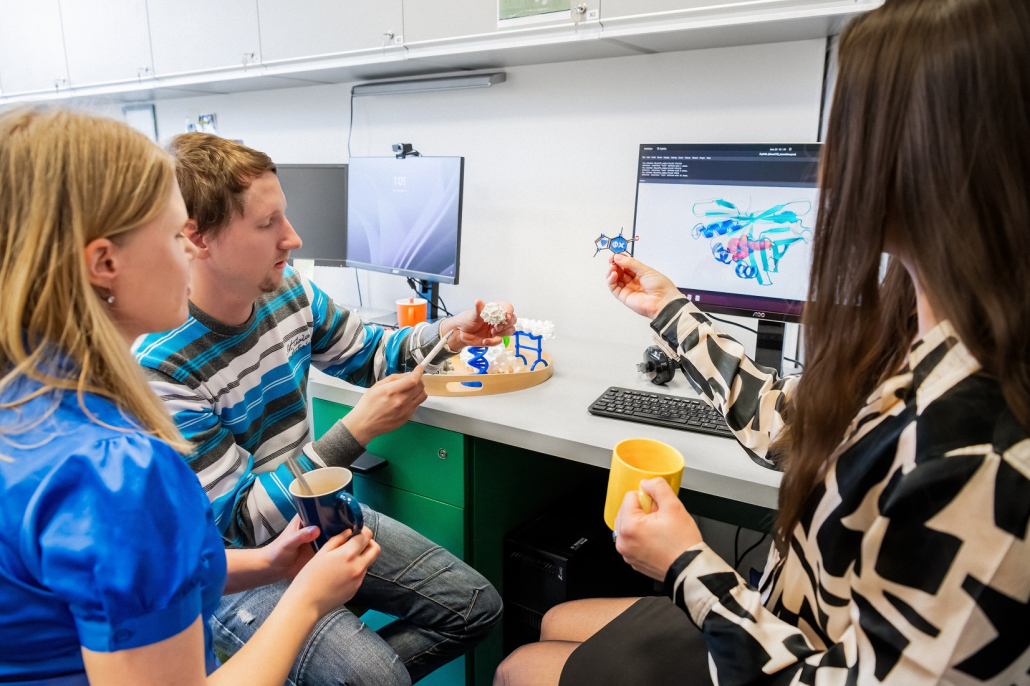
- The research group focuses on protein and metabolic engineering for biomedicine. The group develops new theoretical concepts, software tools and lab-on-chip technologies for protein engineering. It uses these newly developed tools for the design of proteins with improved properties for biocatalysis, biodegradation, biosensing, cell culturing and differentiation. The group has published more than 200 original articles, 20 book chapters and filed 7 international patents and founded the biotechnology spin-off Enantis Ltd.
- Bioinformatics – identification of interesting genes in genomic databases for molecular cloning and experimental characterization
- Lab-on-Chip Technologies – development of microfluidic lab-on-chip technologies for biochemical and biomedical research
- Pathway Engineering – design and construction of bacterial strains expressing newly assembled biochemical pathways
- Protein Stabilization – computational design of stabilization mutations
- International Collaborations: ETH Zurich, Switzerland; Tohoku University, Katahira, Japan; University of Cambridge, Cambridge, UK; Novo Nordisk Foundation Center for Biosustainability (CFB), Copenhagen, Denmark; Spanish National Research Council (CSIC), Madrid, Spain
Main goals
- Computational design and engineering of hyperstable proteins
- Application of novel theoretical concepts for proteins engineering
- Development of user-friendly software tools and microfluidic chips
Technological equipment
- Computer cluster
- Circular dichroism
- Colony picking robot
- Hardware and software for computer modeling
- Microfluidics
- Pipetting robot
- Rapid quench-flow system
- Stopped flow-kinetic system
Team Members:
- prof. RNDr. Zbyněk Prokop, Ph.D. – Principal Investigator
- Mgr. David Bednář, Ph.D. – Senior IT Specialist
- Ing. Simeon Borko – Application Developer
- Mgr. Martina Damborská – Research Coordinator
- prof. Dr. Mgr. Jiří Damborský – Senior Research Team Leader
- Saltuk Mustafa Eyrilmez, Ph.D. – Postdoctoral Researcher
- Irena Halíková – Lab Technician
- Ing. Tomáš Henek – Junior Researcher
- prof. Ing. Lenka Hernychová, Ph.D. – Senior Postdoctoral Researcher
- Petr Kabourek – Senior Application Developer
- Mgr. David Kovář, Ph.D. – Senior IT Specialist
- Mgr. Josef Kučera, Ph.D. – Postdoctoral Researcher
- Ing. Dávid Lacko – Application Developer
- Ing. Marika Majerová– Lab Technician
- RNDr. Ing. Martin Marek, Ph.D., MBA – Postdoctoral Researcher
- Sérgio Manuel Marques, Ph.D. – Postdoctoral Researcher, Research Specialist
- Stanislav Mazurenko, Ph.D. – Senior Postdoctoral Researcher
- MUDr, Jan Mičan – Research Specialist
- Ing. Miloš Musil, Ph.D. – Application Developer
- Mgr. Šárka Nevolová, Ph.D. – Research Coordinator
- Joan Planas Iglesias, Ph.D. – Senior Application Developer
- Ing. Monika Rosinská – Application Developer
- Mgr. Karolina Sedláčková – Lab Specialist
- Mgr. Adam Paulin Urminský – Ph.D. Student
- Mgr. Pavel Vaňáček, Ph.D. – Postdoctoral Researcher
- Mgr. Michal Vašina, Ph.D. – Junior Researcher
- Naina Verma, MSc – Ph.D. Student